
ECUST Achieved Major Breakthrough in AI-Empowered Enzymatic Synthesis of Long Single-Stranded DNA
The research team led by Professor Haizhen Wu from the Pathway Engineering Laboratory, School of Biotechnology, ECUST, has made a significant leap in the interdisciplinary field of artificial intelligence and enzyme engineering. The paper, Machine Learning-Optimized Long Single-Stranded DNA Synthesis Technology Empowers High-Precision Diagnostic-Therapeutic Integration in Living Cells, was recently published in the journal Nucleic Acids Research.
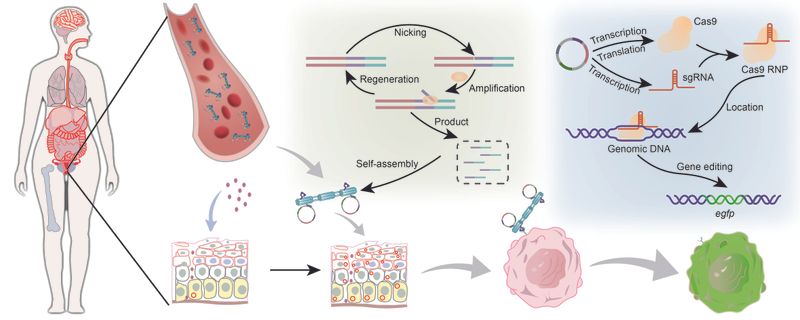
Long single-stranded DNA (ssDNA) is a critical material for constructing DNA nanostructures, gene therapy vectors, and molecular diagnostic probes. However, the synthesis of high-purity ssDNA exceeding 10,000 bases has long been remained an internationally recognized technical bottleneck. To overcome this, the team developed an intelligent, enzyme-assisted replication platform named “Ouroborosyn-ssDNA”. By utilizing machine learning algorithms, the platform globally optimized multiple reaction parameters, including temperature, ion concentrations, and the ratios of key enzymes (phi29 DNA polymerase and Nb.BbvCI nicking enzyme).
This approach enabled the team to systematically reveal, for the first time, the critical mechanism through which magnesium ion kinetics and temperature synergistically regulate the processivity of phi29 DNA polymerase. Within an optimized format buffer system, this technology successfully achieved the large-scale synthesis of high-purity ssDNA up to 15,000 nucleotides in length. Notably, the yield is 4.73 times that of comparable commercial methods, fully meeting clinical application standards.
By pioneering the interdisciplinary path of Machine Learning-Optimized Biocatalysis, this research achieves comprehensive breakthroughs spanning rational enzyme reaction design, intelligent synthesis process optimization, and high-end biomedical applications. The high-efficiency long-strand ssDNA preparation technology established in this study provides essential material support for frontier fundamental research, such as DNA nanomachines and molecular computers.
Furthermore, it demonstrates broad application potential in biomedical fields, including gene therapy, novel vaccine development, and molecular diagnostics. Particularly in the current era of rapid advancement in mRNA vaccines and genomic medicine, this technology is poised to significantly accelerate the research and development pipeline for next-generation therapeutic drugs.
Yue Zhang, a 2025 master’s graduate, is the first author of the paper. Pengfei Li and Professor Haizhen Wu serve as co-corresponding authors, with guidance from Professor Huizhan Zhang.